The title of this piece shouldn’t shock anyone who has taken an introductory statistics course. Statistics is full of terms that have a specific statistical meaning apart from their everyday meaning. A few examples:
Significant, confidence, power, random, mean, normal, credible, moment, bias, interaction, likelihood, error, loadings, weights, hazard, risk, bootstrap, information, jack-knife, kernel, reliable, validity; and that’s just the tip of the iceberg. (Of course, one’s list gets bigger the more statistics courses one takes.)
It should come as no surprise that the general public mistakes a term’s statistical meaning for its general english meaning when nearly every word has some sort of dual-meaning.
Philip Tromovitch (2015) has recently put out a neat paper in which he surveyed a little over 1,000 members of the general public on their understanding of the meaning of “significant,” a term which has a very precise statistical definition: assuming the null hypothesis is true (usually defined as no effect), discrepancies as large or larger than this result would be so rare that we should act as if the null hypothesis isn’t true and we won’t often be wrong.
However, in everyday english, something that is significant means that it is noteworthy or worth our attention. Rather than give a cliched dictionary definition, I asked my mother what she thought. She says she would interpret a phrase such as, “there was a significant drop in sales from 2013 to 2014” to indicate that the drop in sales was “pretty big, like quite important.” (thanks mom 🙂 ) But that’s only one person. What did Tromovitch’s survey respondents think?
Tromovitch surveyed a total of 1103 people. He asked 611 of his respondents to answer this multiple choice question, and the rest answered a variant as an open ended question. Here is the multiple choice question to his survey respondents:
When scientists declare that the finding in their study is “significant,” which of the following do you suspect is closest to what they are saying:
- the finding is large
- the finding is important
- the finding is different than would be expected by chance
- the finding was unexpected
- the finding is highly accurate
- the finding is based on a large sample of data
Respondents choosing the first two responses were considered to be incorrectly using general english, choosing the third answer was considered correct, and choosing any of the final three were considered other incorrect answer. He separated general public responses from those with doctorate degrees (n=15), but he didn’t get any information on what topic their degree was in, so I’ll just refer to the rest of the sample’s results from here on since the doctorate sample should really be taken with a grain of salt.
Roughly 50% of respondents gave a general english interpretation of the “significant” results (options 1 or 2), roughly 40% chose one of the other three wrong responses (options 4, 5, or 6), and less than 10% actually chose the correct answer (option 3). Even if they were totally guessing you’d expect them to get close to 17% correct (1/6), give or take.
But perhaps multiple choice format isn’t the best way to get at this, since the prompt itself provides many answers that sound perfectly reasonable. Tromovitch also asked this as an open-ended question to see what kind of responses people would generate themselves. One variant of the prompt explicitly mentions that he wants to know about statistical significance, while the other simply mentions significance. The exact wording was this:
Scientists sometimes conclude that the finding in their study is “[statistically] significant.” If you were updating a dictionary of modern American English, how would you define the term “[statistically] significant”?
Did respondents do any better when they can answer freely? Not at all. Neither prompt had a very high success rate; they had correct response rates at roughly 4% and 1%. This translates to literally 12 correct answers out of the total 492 respondents of both prompts combined (including phd responses). Tromovitch includes all of these responses in the appendix so you can read the kinds of answers that were given and considered to be correct.
If you take a look at the responses you’ll see that most of them imply some statement about the probability of one hypothesis or the other being true, which isn’t allowed by the correct definition of statistical significance! For example, one answer coded as correct said, “The likelihood that the result/findings are not due to chance and probably true” is blatantly incorrect. The probability that the results are not due to chance is not what statistical significance tells you at all. Most of the responses coded as “correct” by Tromovitch are quite vague, so it’s not clear that even those correct responders have a good handle on the concept. No wonder the general public looks at statistics as if they’re some hand-wavy magic. They don’t get it at all.
My takeaway from this study is the title of this piece: the general public has no idea what statistical significance means. That’s not surprising when you consider that researchers themselves often don’t know what it means! Even professors who teach research methods and statistics get this wrong. Results from Haller & Krauss (2002), building off of Oakes (1986), suggest that it is normal for students, academic researchers, and even methodology instructors to endorse incorrect interpretations of p-values and significance tests. That’s pretty bad. It’s one thing for first-year students or the lay public to be confused, but educated academics and methodology instructors too? If you don’t buy the survey results, open up any journal issue in any psychology journal and you’ll find tons of examples of misinterpretation and confusion.
Recently Hoekstra, Morey, Rouder, & Wagenmakers (2014) demonstrated that confidence intervals are similarly misinterpreted by researchers, despite recent calls (Cumming, 2014) to totally abandon significance tests in lieu of confidence intervals. Perhaps we could toss out the whole lot and start over with something that actually makes sense? Maybe we could try teaching something that people can actually understand?
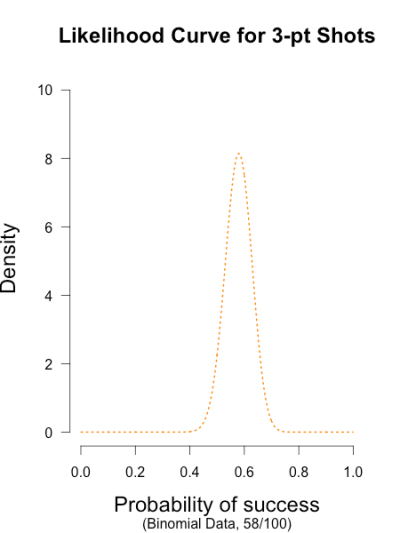
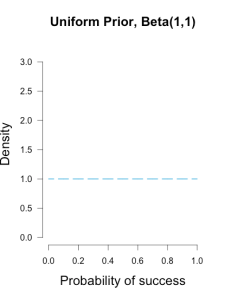
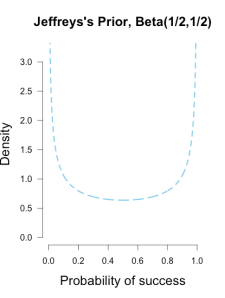
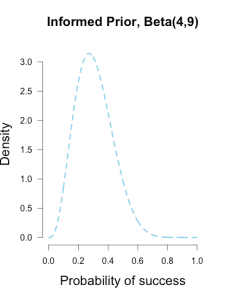
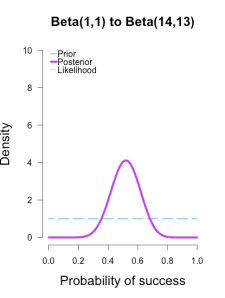
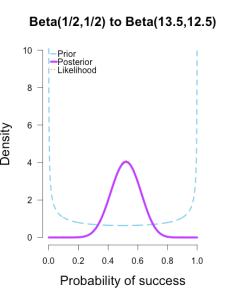
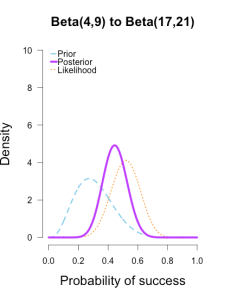
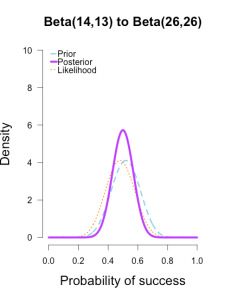
I’ve heard of this cool thing called Bayesian statistics we could try.
References
Cumming, G. (2014). The new statistics: Why and how. Psychological Science, 25(1), 7-29.
Haller, H., & Krauss, S. (2002). Misinterpretations of significance: A problem students share with their teachers. Methods of Psychological Research, 7(1), 1-20.
Hoekstra, R., Morey, R. D., Rouder, J. N., & Wagenmakers, E. J. (2014). Robust misinterpretation of confidence intervals. Psychonomic Bulletin & Review, 21(5), 1157-1164.
Oakes, M. W. (1986). Statistical inference: A commentary for the social and behavioural sciences. Wiley.
Tromovitch, P. (2015). The lay public’s misinterpretation of the meaning of ‘significant’: A call for simple yet significant changes in scientific reporting. Journal of Research Practice, 1(1), 1.